Fuzz testing molecular representation using deep variational anomaly generation.
NOGUEIRA, Victor Henrique Rabesquine; SHARMA, Rishabh; GUIDO, Rafael Victorio Carvalho; KEISER, Michael James.
NOGUEIRA, Victor Henrique Rabesquine; SHARMA, Rishabh; GUIDO, Rafael Victorio Carvalho; KEISER, Michael James.





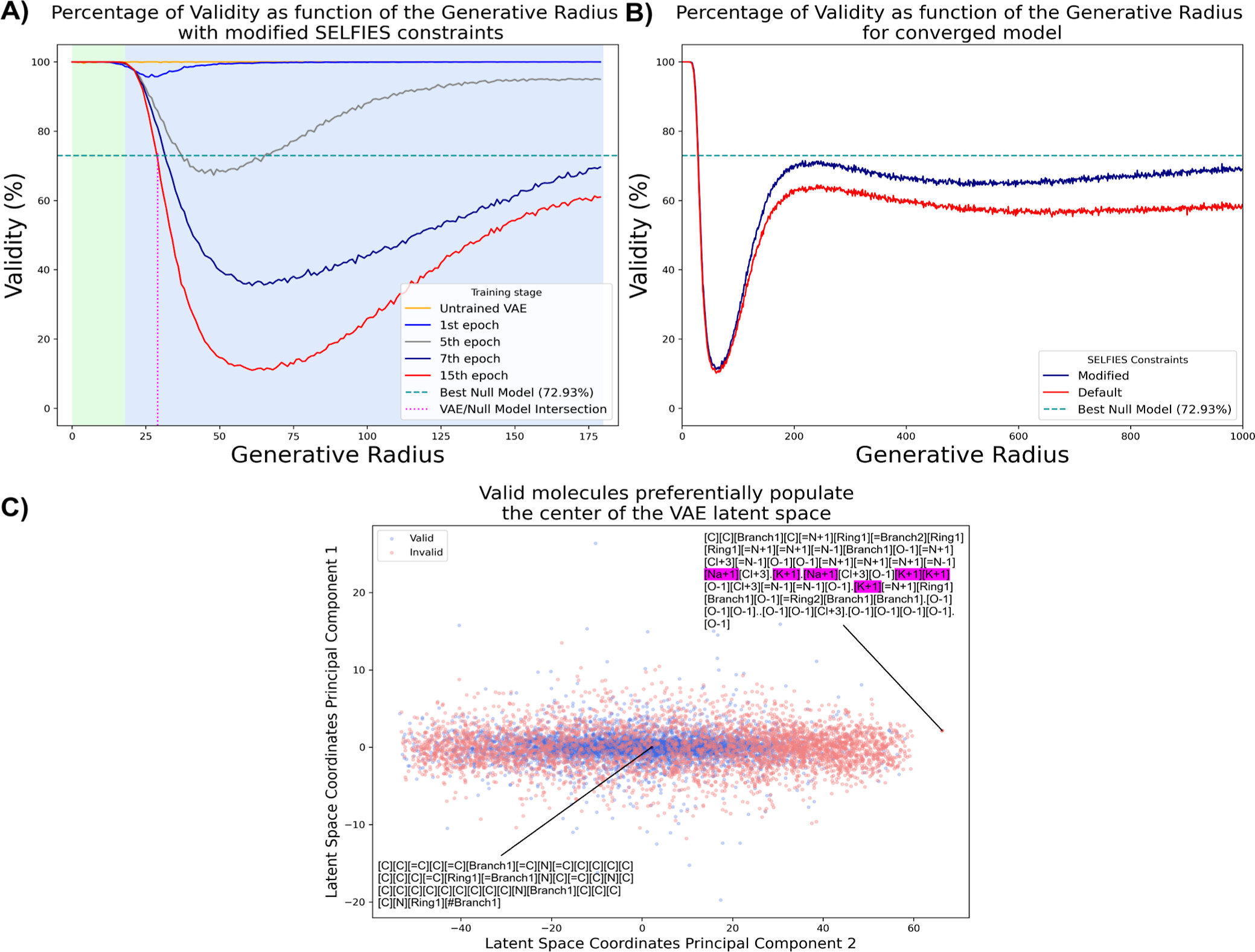 Abstract: Researchers are developing increasingly robust molecular representations, motivating the need for thorough methods to stress-test and validate them. Here, we use a variational auto-encoder (VAE), an unsupervised deep learning model, to generate anomalous examples of SELF-referencIng Embedded Strings (SELFIES), a popular molecular string format. These anomalies defy the assertion that all SELFIES convert into valid SMILES strings. Interestingly, we find specific regions within the VAE?s internal landscape (latent space), whose decoding frequently generates inconvertible SELFIES anomalies. The model?s internal landscape self-organization helps with exploring factors affecting molecular representation reliability. We show how VAEs and similar anomaly generation methods can empirically stress-test molecular representation robustness. Additionally, we investigate reasons for the invalidity of some discovered SELFIES strings (version 2.1.1) and suggest changes to improve them, aiming to spark ongoing molecular representation improvement.
Abstract: Researchers are developing increasingly robust molecular representations, motivating the need for thorough methods to stress-test and validate them. Here, we use a variational auto-encoder (VAE), an unsupervised deep learning model, to generate anomalous examples of SELF-referencIng Embedded Strings (SELFIES), a popular molecular string format. These anomalies defy the assertion that all SELFIES convert into valid SMILES strings. Interestingly, we find specific regions within the VAE?s internal landscape (latent space), whose decoding frequently generates inconvertible SELFIES anomalies. The model?s internal landscape self-organization helps with exploring factors affecting molecular representation reliability. We show how VAEs and similar anomaly generation methods can empirically stress-test molecular representation robustness. Additionally, we investigate reasons for the invalidity of some discovered SELFIES strings (version 2.1.1) and suggest changes to improve them, aiming to spark ongoing molecular representation improvement. @article={003236491,author = {NOGUEIRA, Victor Henrique Rabesquine; SHARMA, Rishabh; GUIDO, Rafael Victorio Carvalho; KEISER, Michael James.},title={Fuzz testing molecular representation using deep variational anomaly generation},journal={Journal of Chemical Information and Modeling},note={v. 65, n. 4, p. 1911-1927 + supporting information},year={2025}}
@article={003236491,author = {NOGUEIRA, Victor Henrique Rabesquine; SHARMA, Rishabh; GUIDO, Rafael Victorio Carvalho; KEISER, Michael James.},title={Fuzz testing molecular representation using deep variational anomaly generation},journal={Journal of Chemical Information and Modeling},note={v. 65, n. 4, p. 1911-1927 + supporting information},year={2025}}